Research
Published 10 June 2019Has Intensive-Fishing Shaped Snapper Evolution?
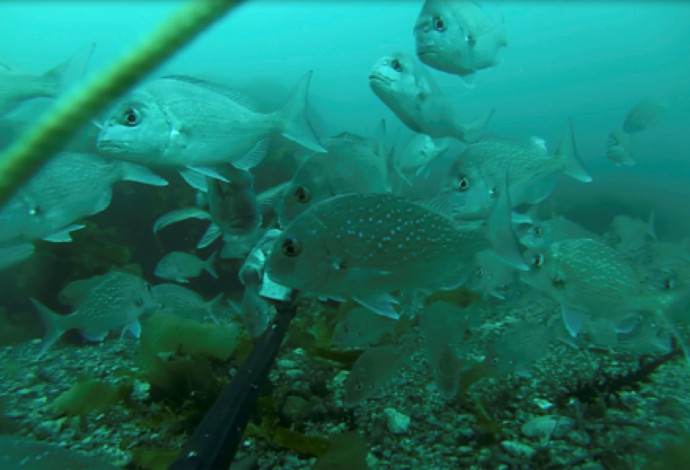

Snapper (Chrysophrys auratus; above) is one of Aotearoa New Zealand’s most important fisheries species and has been an important source of kai moana for Māori since their arrival ca. 700 years ago. With European arrival, these stocks became so exploited that snapper was the first fishery to be regulated in the late 19th century with a minimum catch size. With the introduction of commercial fisheries and power-driven net trawlers, by the 1980s snapper stocks were nearing collapse. The introduction of new fisheries management control systems has helped stop further declines, but snapper still experience high catch levels by both recreational and commercial fishers.
Figure 1: Size-frequency histograms of snapper from archeological collections (above) show that fish were larger when compared to sizes found in the modern fishery (below).
Intense size-selective fishing has been linked to reductions in average fish size and early maturation (Figure 1). However, it is unknown if these effects are irreversible and if they are caused by genetic changes within snapper populations. Understanding how industrial-scale fishing affects fish stocks is not only a fascinating evolutionary question, but it is important to the long term sustainability of these fisheries, as well as marine conservation efforts. PI Dr Peter Ritchie (Victoria University of Wellington) leads the Marsden Funded team to address these questions.

Figure 2: Up to 600 years old snapper jaw bones collected from Māori middens from which ancient DNA has been sequenced.
Marsden Funded PhD student Tom Oosting, working in the research group of Dr Ritchie, has been sampling and comparing the genome sequences of contemporary New Zealand snapper to pre-commercially fished populations. Historical snapper bone samples (Figure 2) were derived from archaeological excavations of Māori middens (Te Papa museum) dating back to 600 years ago. The relatively short human occupation of New Zealand provides a unique experimental situation enabling this project. In many other parts of the world large-scale human exploitation began thousands of years ago, putting the hope of sampling the baseline (pre-human exploitation) population out of reach for most fisheries species. The ancient DNA samples from snapper bones derived from Māori middens represent the genetic composition of near-pristine snapper populations prior to industrial-level exploitation. This group has now sequenced the genomes of snapper up to 600 years old, utilising specialised ancient DNA labs run by Dr Nic Rawlence (Otago University), and Dr Bastiaan Star and Associate Professor Sanne Boessenkool (Oslo University).
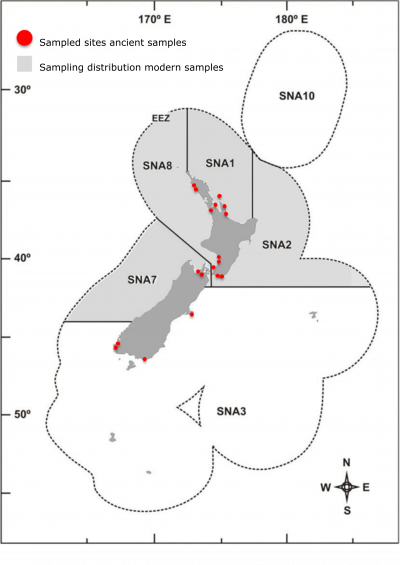
Figure 3: Distribution of historical sample sites and modern fishery ranges for snapper included in this study.
This group have also collected over a thousand contemporary snapper samples that span the entire range of the New Zealand fishery (Figure 3). Sampling was made possible through the assistance of surveys conducted by the Ministry of Primary Industries (MPI), the National Institute of Water and Atmospheric research (NIWA), and with the help of recreational fishing competitions run by the Tatapouri fishing club, the Hawke’s Bay sports fishing club, and the Petone working men’s club. All samples were collected from individual snapper that were caught for commercial/recreational consumption or under other scientific studies insuring no snapper were caught and killed specifically for this study.
Technological advances in genome sequencing allow researchers to obtain entire genome sequences, from both archeological and comtemporary samples. Work is ongoing to build the bioinformatic pipeline allowing comparison of hundreds of ancient and contemporary snapper genomes to the snapper reference genome assembled by team member Dr Maren Wellenreuther (Plant and Food Research, Auckland University). This will allow them to identify thousands of informative DNA bases that show variation both between individual snapper and between ancient and contemporary snapper populations. This information will then be used to identify snapper stock structure, demographic changes over time, and snapper evolutionary change. Already they are starting to identify novel differences among snapper and patterns of genetic structure. In the near future, they plan to compare the genomes from historical and contemporary samples in order to assess the extent of effects due to human exploitation of a commercially caught species.
RESEARCHER
Dr Peter Ritchie and Tom Oosting (PhD Candidate)
ORGANISATION
Victoria University of Wellington
FUNDING SUPPORT
Marsden Fund
CONTRACT OR PROJECT ID
VUW1602
